Setting up the Persephone System
First of all, you are welcome to use the free Persephone instance at https://web.persephonesoft.com. This is our web portal that runs Persephone with many popular genomes. The fully-functional application will work in your web browser with the genomes loaded into our database. It will also allow you to add your own data sets (up to a 5 GB disk quota). If this is all you need, please feel free to keep using the website and give us your feedback. If you prefer to work with your proprietary data, then yes, you will need to set up your own instance of Persephone. Please contact our support and, once we agree on the licensing terms, we will create an FTP account for you to download the software and the license. We will guide you through all the necessary steps of installation and learning the system.
This section provides the steps and guidelines needed to install and configure your own Persephone system.
Note
The Persephone System Setup Guide is intended for administrators only.
Note
We highly recommend installing the Persephone software stack by using a single Docker image. Once the Docker image is installed, you can start using PersephoneShell to populate the data and run the Persephone web client. If you prefer to get familiar with the Persephone architecture, please keep reading the text below.
Logical System Diagram
The following figure shows a logical (conceptual) representation of the Persephone system.
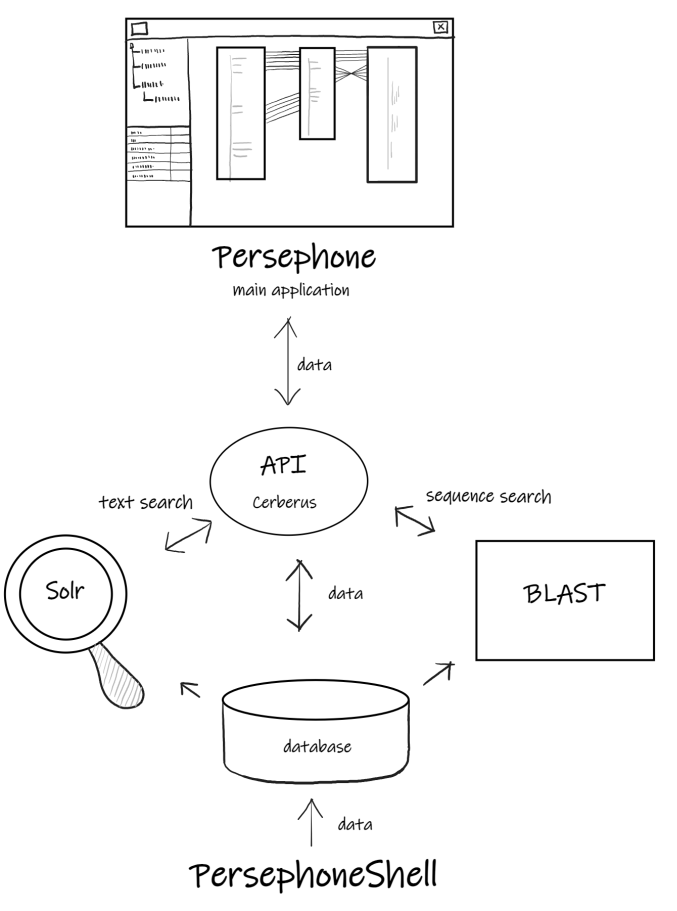
The majority of the data is stored in the database. Users can drag&drop external files onto the client application or reference them by URL to create their private tracks or entire map sets (genomes).
The main components
The required components of the Persephone system are: a database, an API server, a Solr server, and the loader application PersephoneShell. The majority of the genomic data is stored in the database. The API server reads the data and sends it to the client application responding to its requests. The Apache Solr server provides the fast search services. PersephoneShell loads the data to the database checking for its consistency.
The back end (Note: the Docker image has all the components preinstalled) |
|
The main application |
|
Installation steps without details (when not using the Docker image)
1. Install the database server, create an empty database and a new database user.
2. Install Solr search engine (requires Java). Create a new core.
3. If hosting on linux, install Mono (we need it to run .NET applications).
4. Unpack PersephoneShell from our archive and update configuration values: the database connection string, path for BLAST binaries, URL to Solr, location of external files, etc.
5. Unpack WebCerberus from our archive and modify the configuration: the database connection string, path for BLAST binaries and index files, URL to Solr, etc.
The advanced security configuration steps to enable user registration are described here.
Just to remind you, we provide a Persephone Docker image that already has all the needed components pre-installed and configured. Just spin the container and navigate to a URL to see the live application.
System Requirements
The web client of Persephone runs on any desktop OS, as long as it provides a modern web browser. The recommended hardware for the client machine would look like this:
Client:
|
Processor Type |
Dual core |
|
Processor Speed |
2.8 GHz |
|
Memory |
8 GB minimum |
|
Local Storage |
1 GB for program files and the data cache. |
Server:
The requirements for the server side are higher (depending on the load, for small teams it can run on a regular laptop).
|
Processor Type |
Quad core |
|
Processor Speed |
2.8 GHz |
|
Memory |
16 GB minimum |
|
Local Storage |
100 GB for program files and the data cache. The data files can take more. |
While most of the data is stored in the database, the bulky entries, such as genomic sequences, BAM tracks, or variant binary data, are stored outside the database, in the file system. Such organization reduces stress on the database and dramatically improves the timing of the backup process. There is no need to "export" massive data records from the database in the form of SQL text, which is very time-consuming - the compressed binary files are ready for backup without additional preparation.
