Adding Functional Annotation
Step 6: Adding functional annotation
The last step in our exercise is adding functional annotation by finding matches to SwissProt and saving the information as gene qualifiers. Run the command
create function <mapSetId>
for each map set. For now, this can be done by executing this command in a loop from the linux command prompt. PersephoneShell will be started in batch mode in this format:
psh [psh_options] psh_command [command_options]
For the range of MapSetIds 513..549, we can run the following command using the for command in linux:
for a in {513..549}; do psh create function $a; done
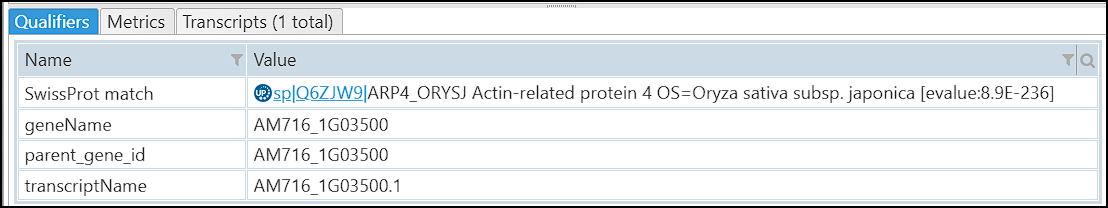
For each of the map sets addressed by MapSetId the command will prepare a FASTA file with protein sequences by either unpacking the corresponding BLASTP sequence collection or by generating the proteins from the database. After that, the protein aligner (most likely, DIAMOND) will be called and its output will be used to produce the qualifier SwissProt match: