Adding Variants
Step 5: Adding genetic variants
The genetic variation data is available in the form of a VCF file at http://cucurbitgenomics.org/v2/ftp/pan-genome/cucumber/variant/cucumber_SNP_446.vcf.gz.
We will load all genotyping samples on all chromosomes, so our INI file for the command add variant is quite simple:
cucumberSNP.ini
[ProcessRun]
[MapSet]
MapSetPath=/Cucumis sativus (pangenome)/Cucumber WI7633
[Variant]
Sources=http://cucurbitgenomics.org/v2/ftp/pan-genome/cucumber/variant/cucumber_SV_446.vcf.gz
[MapMapping]
Here, we are trying to use the VCF file directly from the remote location. In case of slow network, it is recommended to download the file first and reference its local copy to minimize the network traffic during test and loading procedure.
add variant -c $DATA/cucumber/cucumberSNP.ini -v
After loading, the variant data will be visible in the corresponding interfaces of Persephone.
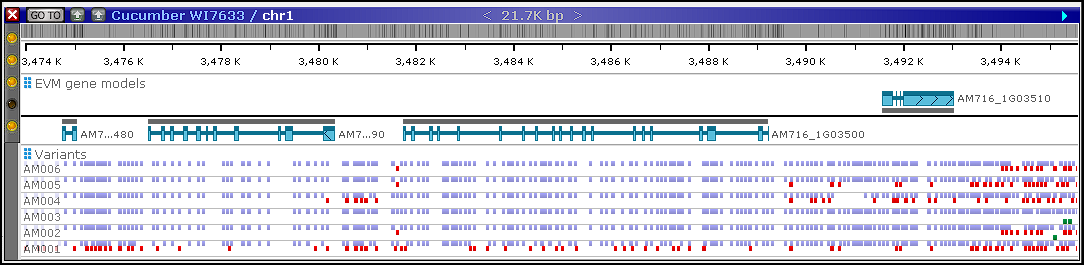
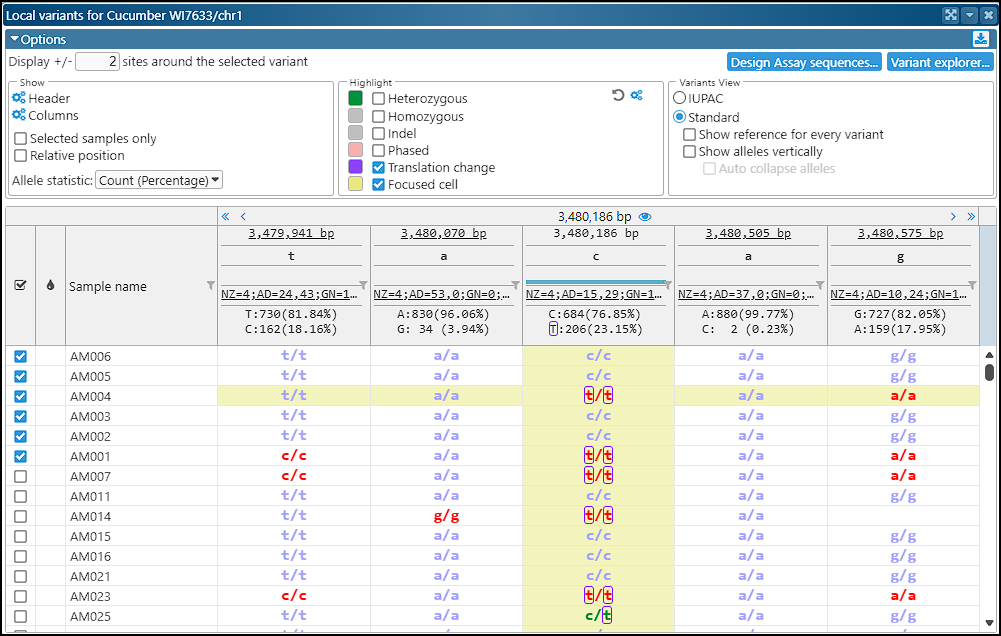
Step 6. Adding functional annotation
