Creating Orthologs
Step 4: Creating orthologs
Loading the gene annotation in the previous step enables whole‑pangenome alignment by linking orthologous gene pairs across all genomes. To accomplish this, we must run create ortholog for every genome pair. With 37 protein sets, this produces 703 genome combinations, including self‑comparisons for paralog detection, and yields roughly 20 million orthologous relationships.
Instead of running the command manually for each pair, we can issue a single create ortholog command that accepts a range of MapSetIds. The system will automatically compute orthologs for every genome combination—both cross‑genome orthologs and within‑genome paralogs—covering the entire pangenome in one operation.
To find which map sets to include into the analysis, we will query MapSetIds for genomes that have 'cucumber' in their name:
PS> list mapset -p cucumber*
513: Cucumber 9110gt 514: Cucumber Chinese Long 9930 v4
515: Cucumber WI2757 516: Cucumber WI7012
517: Cucumber WI7037 518: Cucumber True Lemon
519: Cucumber WI7150 520: Cucumber WI7167
521: Cucumber WI7204 522: Cucumber Poinsett 76
523: Cucumber Marketmore 76 524: Cucumber PI 179678
525: Cucumber PI 197088 526: Cucumber PI 215589
527: Cucumber PI 249561 528: Cucumber PI 330628
529: Cucumber WI5551 530: Cucumber WI7180
531: Cucumber PI 109483 532: Cucumber PI 214155
533: Cucumber PI 462369 534: Cucumber WI7439
535: Cucumber WI7724 536: Cucumber WI7646
537: Cucumber WI7633 538: Cucumber PI 531313A
539: Cucumber WI7651 540: Cucumber WI7698
541: Cucumber WI7773 542: Cucumber PI 183967
543: Cucumber PI 618917 544: Cucumber CG6663
545: Cucumber PI 221440 546: Cucumber PI 500361
547: Cucumber PI 175120 548: Cucumber CG9192
549: Cucumber Cu2
37 mapsets
Our range for MapSetId is 513..549. The command creating the orthologs is simply:
create ortholog 513..549
This task may take a couple of hours.
Now, the maps can be aligned in Persephone:
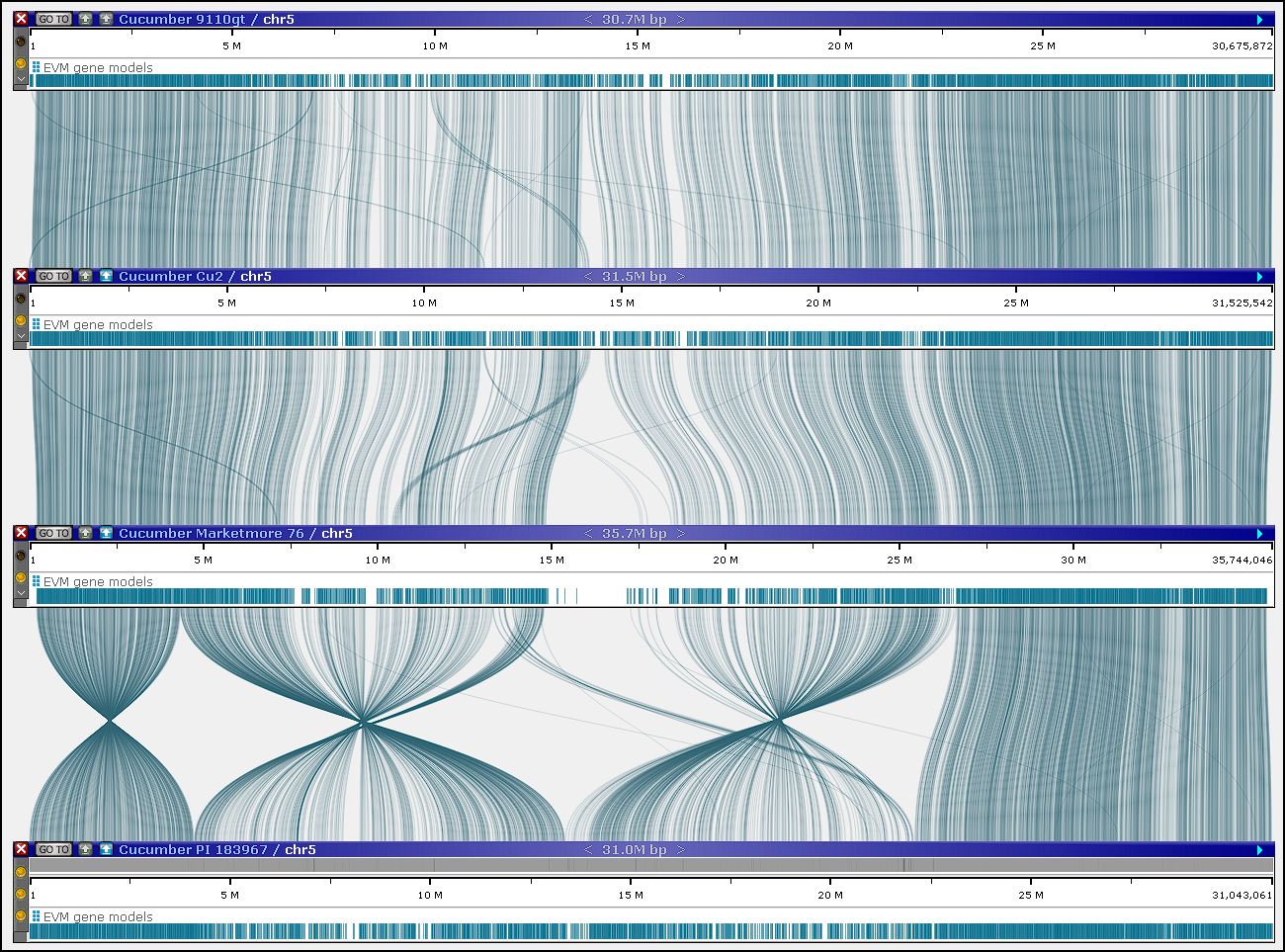
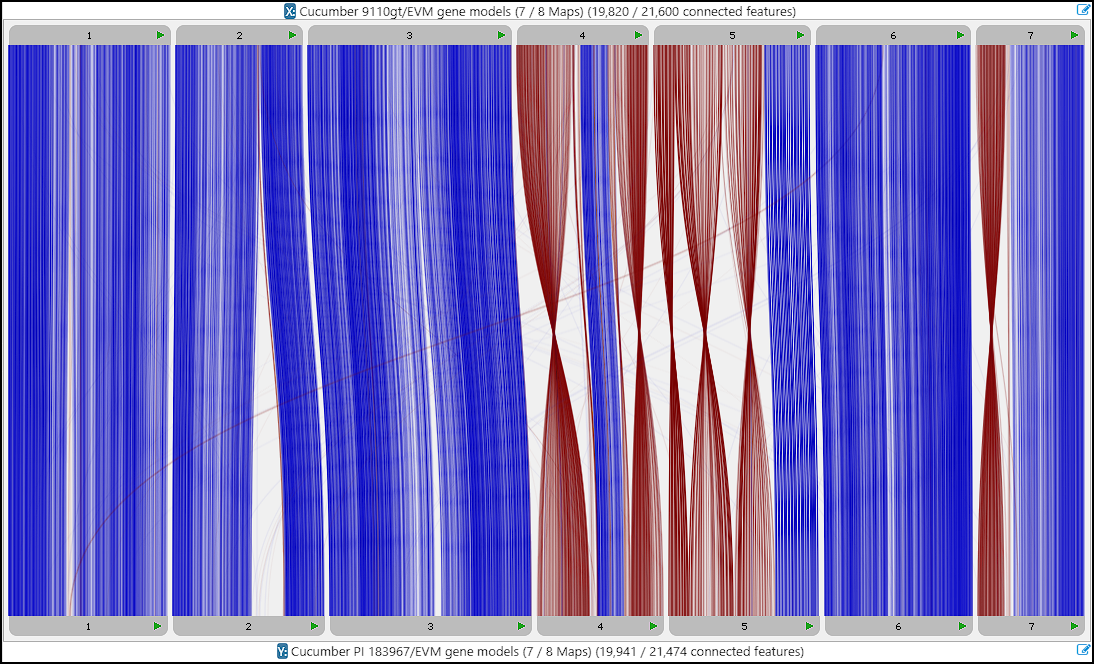
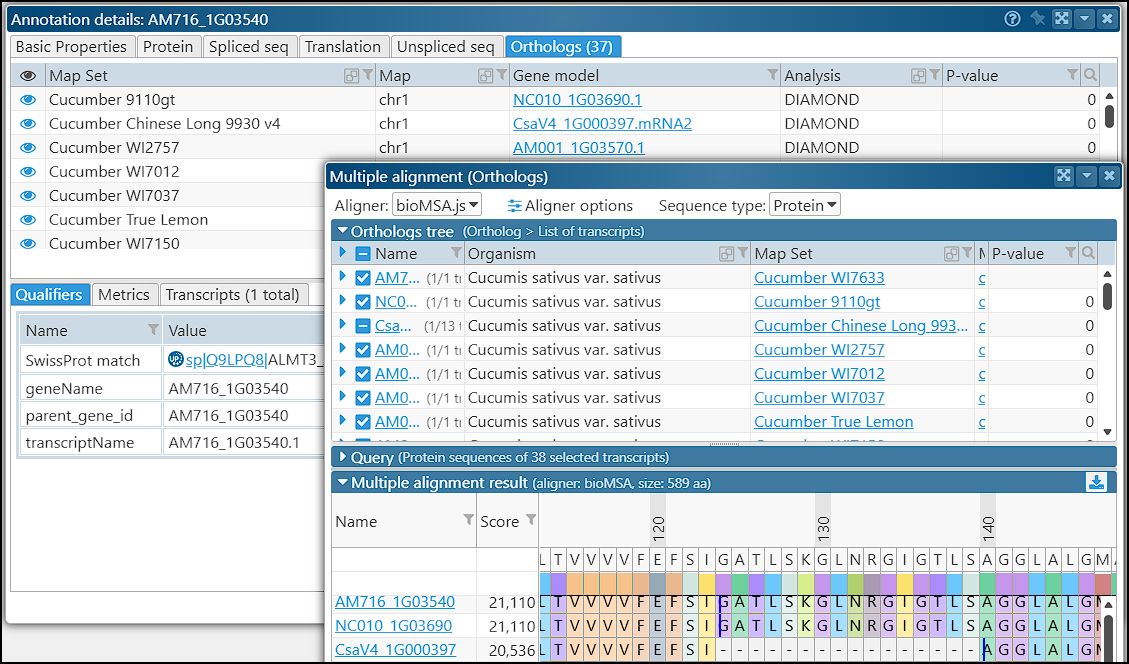
The orthologous proteins for each gene can also be studied in multiple alignment interface available through the gene properties form:
Step 5. Adding variants.
