Markers
A genetic marker is an object that usually has position on some map. A collection of markers can be used to identify individuals or species. Sequences associated with markers can be short DNA sequences surrounding SNPs or long microsatellites. A marker should have a name and a source organism. Besides that, there could be multiple name aliases, sequences and qualifiers. Normally, the markers have positions on one or more maps.
The figure below shows the hierarchy of markers in Persephone.
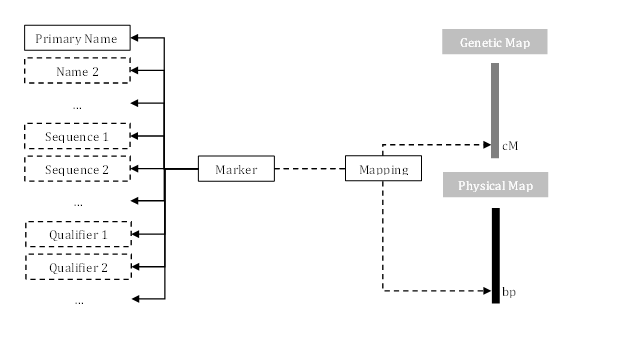
The following sections describe control file sections used to add and modify markers.
Tip
See Add Markers for an example of using a control file to add annotation markers.
MarkerNameType
A single marker can have multiple names. Each name has its own type. For example, a marker named 'adh2' could also be called as 'S45023' or 'X02915' elsewhere. If you want to consider adh2 as 'MARKER_NAME' and other names as 'ALIAS', you define those name types in the [MarkerNameType] section in the INI control file as shown in the example below. If MARKER_NAME and ALIAS already exist, they will be updated; otherwise, added.
The following shows the format for the [MarkerNameType] control file section.
[MarkerNameType]
MARKER_NAME="Primary marker name."
ALIAS="Another name used to call a marker."
In each marker track section, specify types of names and a primary marker name type given file source.
For example, if the adh2 marker is defined in a source GFF file as shown below:
chr1 . SNP 440159 440159 91.68 - . ID=adh2;Name=S45023;Alias=X02915
then Marker section of the control file would be
[Marker]
Tracks=MyMarker1
[MyMarker1]
PrimaryMarkerNameType=MARKER_NAME
GffMarkerNameAttributeKey.ID=MARKER_NAME
GffMarkerNameAttributeKey.Name=ALIAS
GffMarkerNameAttributeKey.Alias=ALIAS
This set of instructions tells that a marker will be created with a primary name MARKER_NAME copied from a GFF attribute ID. There will be other names for this marker of type ALIAS copied from attributes Name and Alias.
TextMarkerName
If the text marker names are defined as follows:
Map_name pos marker_name alias1 alias2
chr1 440159 adh2 S45023 X02915
then the entries in the control file would be:
TextMarkerNameIndex.2=MARKER_NAME
TextMarkerNameIndex.3=ALIAS
Note that the names of keys start with 'Gff' or 'Text' depending on the format of the input source file.
MarkerSequenceType
The following shows the format for the control file MarkerSequenceType section.
[MarkerSequenceType]
SEQ="The full sequence of the marker."
AddMode
Use the AddMode parameter to add repeat markers. The possible values for the AddMode parameter are shown below. Please note, markers can be reused; however, a primary marker name within a map set should be unique (MarkerName, MarkerSequence, MarkerQualifier, Mapping, MappingQualifier).
|
AddMode |
Description |
|
AddAnyway |
Add regardless of duplication. Faster as it does not check. |
|
AddOrDie |
Add if not exists; die (throw exception) otherwise. |
|
AddOrUpdate |
Add if not exists; update otherwise. |
|
AddOrSkip |
Add if not exists; skip otherwise. |
