Web Persephone: Design assay sequences
Persephone's Design assay sequences tool provides a convenient way to generate assay sequences flanking a variant (typically a SNP). You can access the tool in several different ways:
- By selecting Tools | Design assay sequences from the main toolbar; doing so opens a blank Design assay sequences dialog.
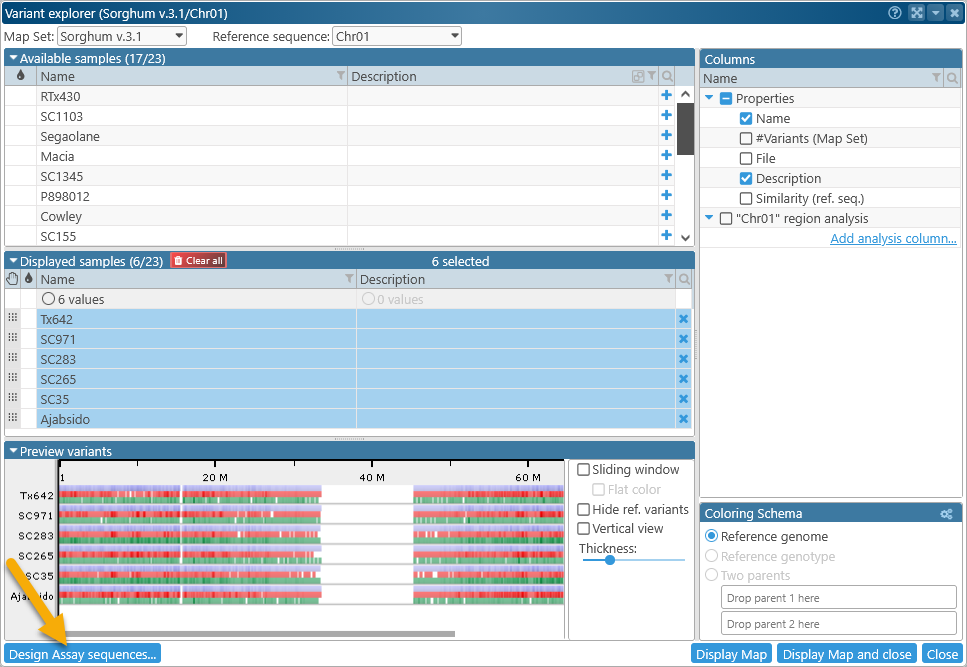
- By clicking the Design assay sequences button in the bottom-left corner of the Variant Explorer dialog. Doing so also opens a blank Design assay sequences dialog, with your current Map Set pre-selected.
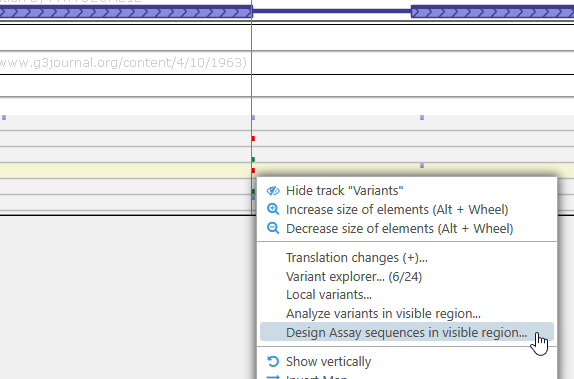
- By right-clicking a variant on the Variants track, and selecting Design assay sequences in visible region from the popup menu. Doing so populates the dialog with anchor sites for each variant in the currently visible region on the map.
The Design assay sequences dialog displays a list of anchor sites at the top, and the resulting assay sequences at the bottom: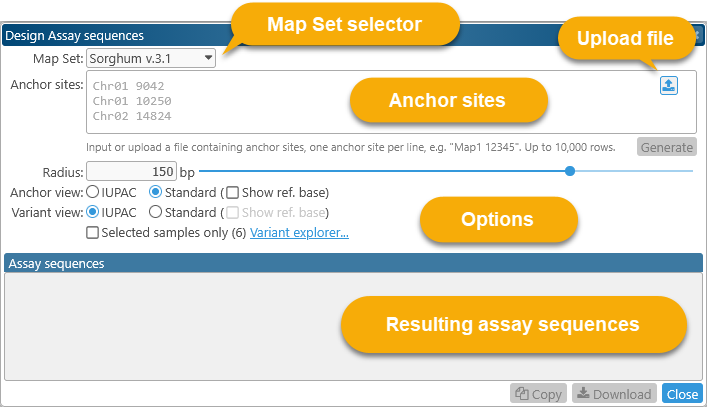
The Map Set selector at the top contains a dropdown list of all Map Sets with available variant samples; this list is identical to the one in the Variant explorer: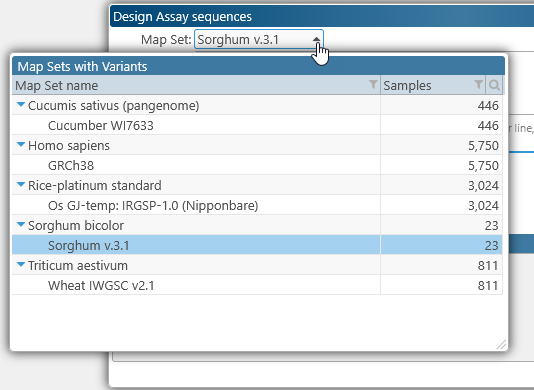
When you select the desired Map Set, the Anchor sites input field will display a prompt with some sample anchor sites for the assay. Each anchor site must consist of two fields, separated by whitespace, similar to "Chr01 1234"; where the first field is the name of the map, and the second field is the location of the anchor site on the map (in nucleotides, 1-based). This location must contain the variant of interest (typically a SNP).
Note
The map names specifying anchor sites are currently case-sensitive. This means that a map name such as "Chr01" will not match a map named "chr01".
You can paste in a list of anchor sites by hand; alternatively, you can click the  button to upload the list from a text file (in the same format). You can also drag-and-drop the file directly onto the input field. The list of anchor sites is currently limited to 10,000 entries.
button to upload the list from a text file (in the same format). You can also drag-and-drop the file directly onto the input field. The list of anchor sites is currently limited to 10,000 entries.
After entering (or uploading) the anchor sites, click the Generate button to generate the assay sequences: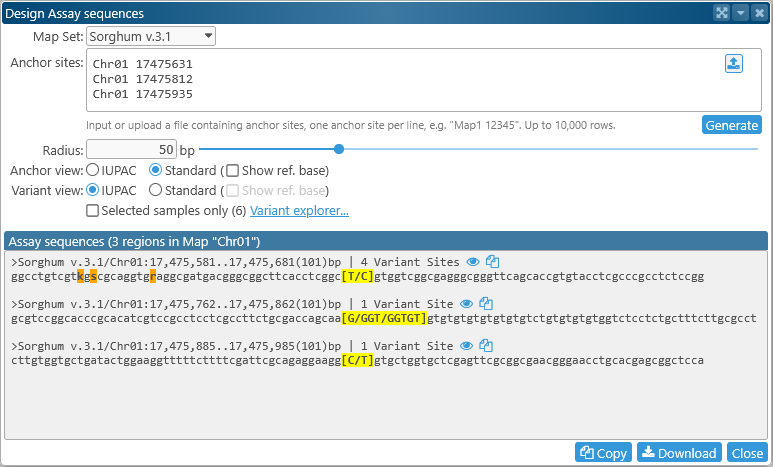
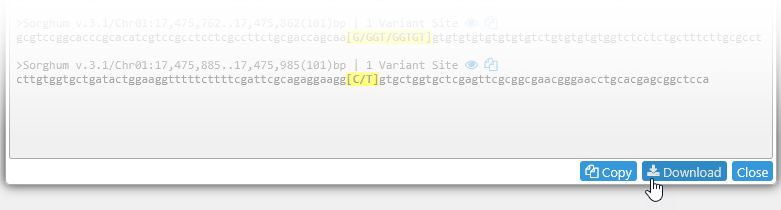
The resulting sequences are generated in FASTA format, with the header for each sequence listing its anchor site, length, and the number of variant sites within the resulting sequence. The variant at each anchor site is highlighted in yellow; you will see a warning if no variant could be found at the site. Flanking variants are highlighted in dark orange; note that the presence of flanking variants may be detrimental for the resulting assay. By default, main variants are displayed in full, and flanking variants are displayed in IUPAC shorthand. You can change this behavior using the controls in the middle of the dialog:
- Radius: Use this slider to set the length of the sequences flanking each anchor site.
- Anchor view / Variant view: Set display options for the variants at anchor sites as well as any flanking variants:
- IUPAC: Uses IUPAC codes to represent variants, e.g. "C/T" is shown as "Y".
- Standard: Displays all allele sequences in full, e.g. "C/T".
- Show ref. base: Displays base in the reference sequence at the variant site, e.g. "C:C/T"
- Selected samples only: By default, all available samples are considered for assay design; check this checkbox to only consider those samples that are currently selected in the Variant explorer. The number (in parentheses) shows the count of samples that are currently selected.
After the assay sequences have been generated, you can click the ![]() button next to each sequence to open the corresponding map, zoom in to the region covered by the sequence, and highlight it:
button next to each sequence to open the corresponding map, zoom in to the region covered by the sequence, and highlight it: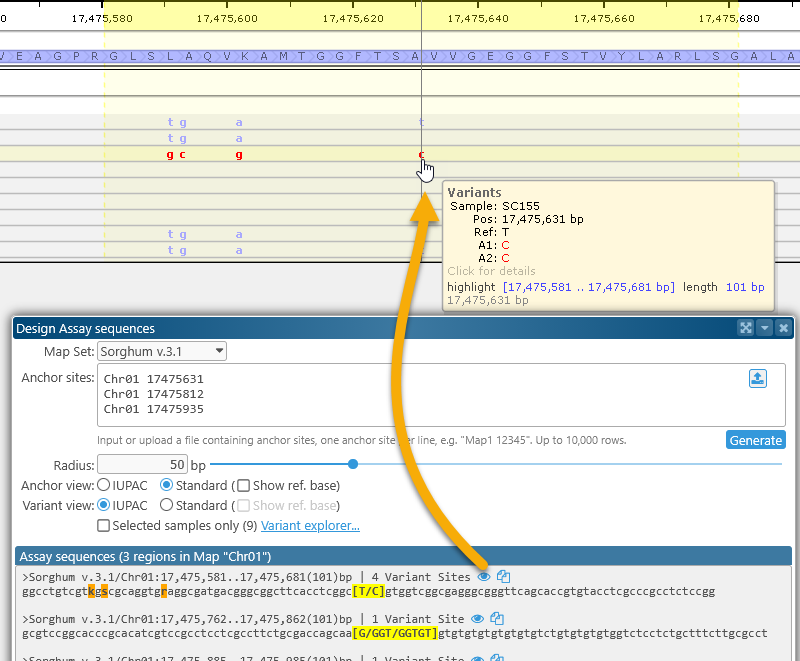
You can also click the  button next to a sequence to copy it to the clipboard.
button next to a sequence to copy it to the clipboard.
Alternatively, click the Copy button at the bottom of the dialog to copy all the assay sequences to the clipboard, or click the Download button to download them as a FASTA file.