Settings: Tracks
Tracks
This tab of the Settings dialog contains general display options affecting all tracks, as well as specific options for individual track types, which are collected into a set of sub-tabs at the bottom of the dialog:
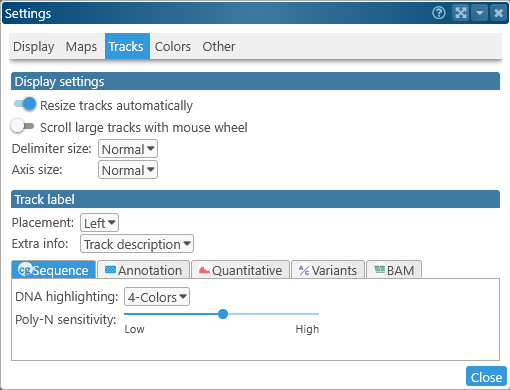
Display settings
- Resize tracks automatically: When this option is turned on, some tracks (such as the Annotation track) will be automatically made wider or narrower depending on the current zoom level, and the number of elements on screen. Turn this option off to always keep tracks at a constant size; this could be useful when making a screenshot for publication.
- Scroll large tracks with mouse wheel: By default, placing the mouse cursor over a map and rolling the mouse wheel zooms the map. Enable this option to instead scroll a large track (e.g. the BAM track) when you roll the mouse wheel over it. If there is nothing to scroll, the mouse wheel will still try to zoom.
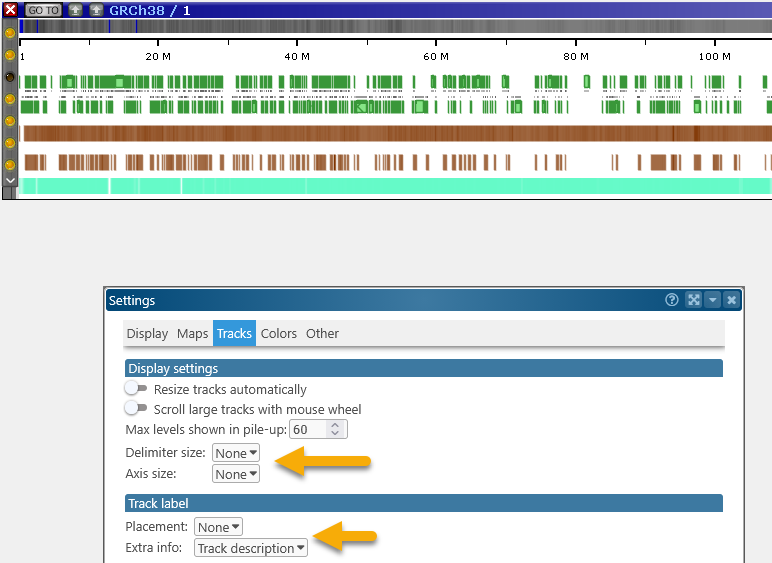
- Delimiter size, Axis size: These options control the thickness of delimiters (gray lines) between tracks, and the axis within tracks:
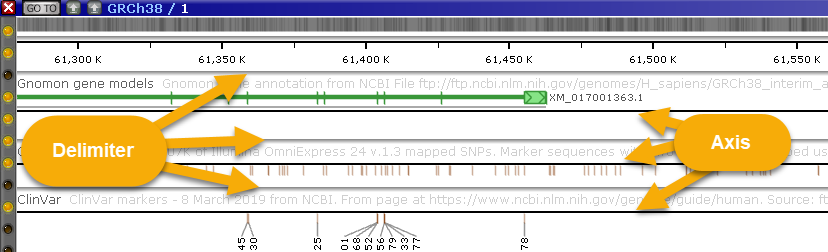
For example, you could set both of these options to None to produce a cleaner output: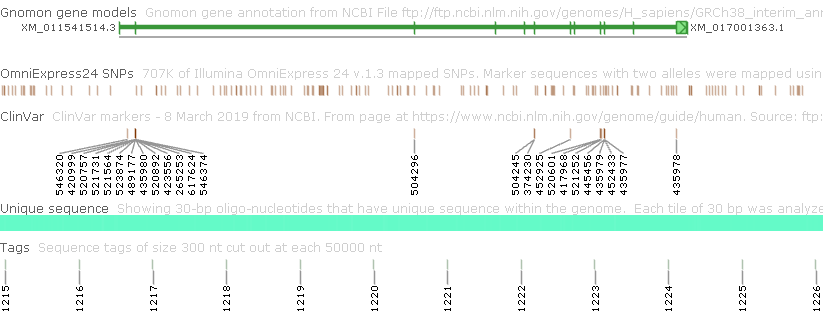
Track label
These options control the labels displaying each track's name and description.
- Placement: Places the track's name on the left side / middle / right side of the map, or turns it off altogether.
- Extra info: Determines what is displayed in the gray description field next to the track's name: the track's description (if any), statistics, or nothing at all.

Turning off both of these labels can be helpful if you wish to save some room on screen: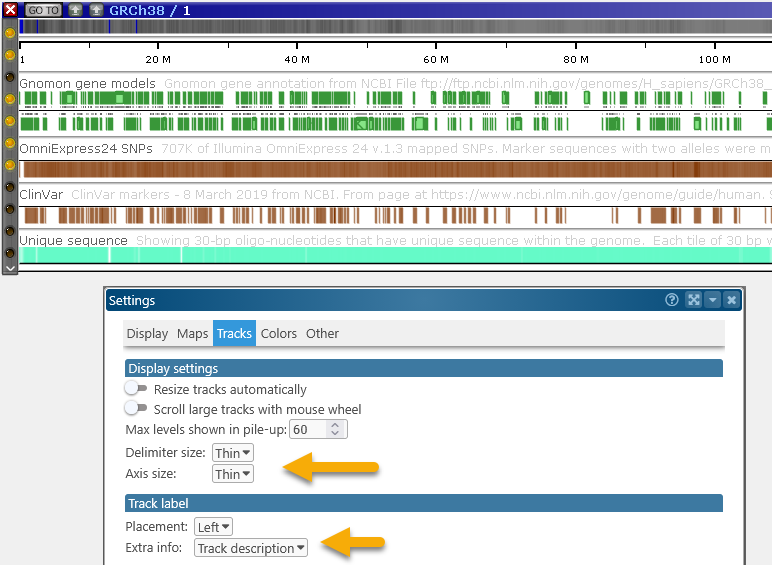
Sequence track
Controls the behavior of the Sequence track, also known as the GC-content track.
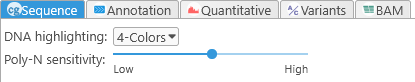
- DNA Highlighting: When the track is zoomed in closely enough to show individual nucleotides, determines how the nucleotides are highlighted: in four distinct colors, grayscale (indicating GC-content), or monochrome.
- Poly-N sensitivity: Adjust this slider to emphasize or de-emphasize poly-N segments; lower values will cause smaller poly-N segments to be hidden when the track is zoomed out.
Annotation track
Controls the appearance of the Annotation track at various zoom levels.
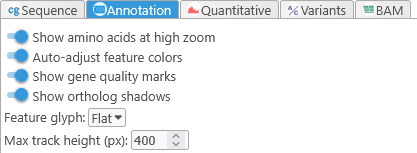
- Show amino acids at high zoom: Shows or hides automatically translated protein sequences for gene annotations:
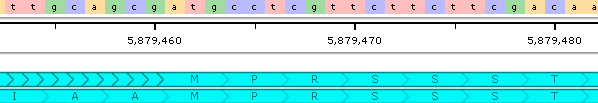
- Auto-adjust feature colors: Automatically adjust the brightness of annotation features to make their exons and CDS indicators easier to read.
- Show gene quality marks: Some gene annotations may possess quality indicators, displayed as colored dots on the upper-left corner of the annotation (ranging from green if the quality is high, to red if the quality is low):
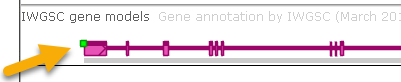
Turn off this option to hide these quality indicators. Note that you can click a gene annotation to view its quality: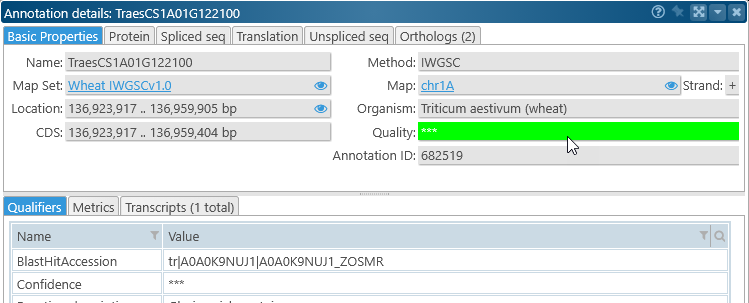
- Show ortholog shadows: Turn off this option to hide "shadows" on the Annotation track's axis that indicate presence (and density) of orthologs:
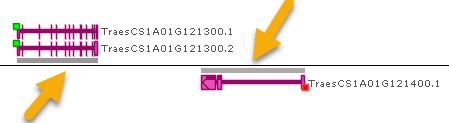
- Feature glyph: Switch between flat and gradient-shaded presentation of gene annotation glyphs. The latter one can be handy to distinguish closely located genes:

- Max track height (px): Sets the maximum height of the track (in pixels). The track will attempt to stay within this size when automatically resizing itself:

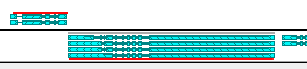
This setting has no effect if you've resized the track manually (by dragging its bottom border).
Quantitative track

- Ruler placement: Use this setting to move the ruler on Quantitative tracks to the left, middle, or right areas of the map; or to turn off the ruler altogether.
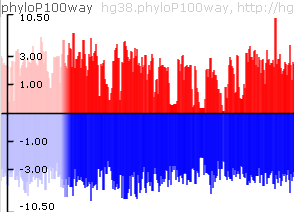
Variants track
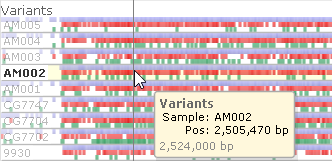
- Sample name opacity: Adjusts the opacity of sample name labels on Variants track. Note that the name of the sample currently under the mouse cursor is always shown fully opaque:
BAM track
- Max track height (px): Sets the maximum height of the track's pile-up section (in pixels). The track will attempt to keep the pile-up within this size when automatically resizing itself:


Note that this setting operates independently of the Max track height for the Annotation track.
